Genomics Technique Accelerate Detection of Foodborne Bacterial Outbreaks
By LabMedica International staff writers
Posted on 15 Dec 2016
Diagnostic testing for foodborne pathogens relies on culture-based techniques that are not rapid enough for real-time disease surveillance and do not give a quantitative picture of pathogen abundance or the response of the natural microbiome.Posted on 15 Dec 2016
Metagenomics identifies the microbes present by sequencing the entire DNA present in a sample and comparing the genomic data to a database of known microbes. In addition to identifying the bacteria present in the samples, the methodology can also measure the relative abundance of each microbial species and their virulence potential, among other things.
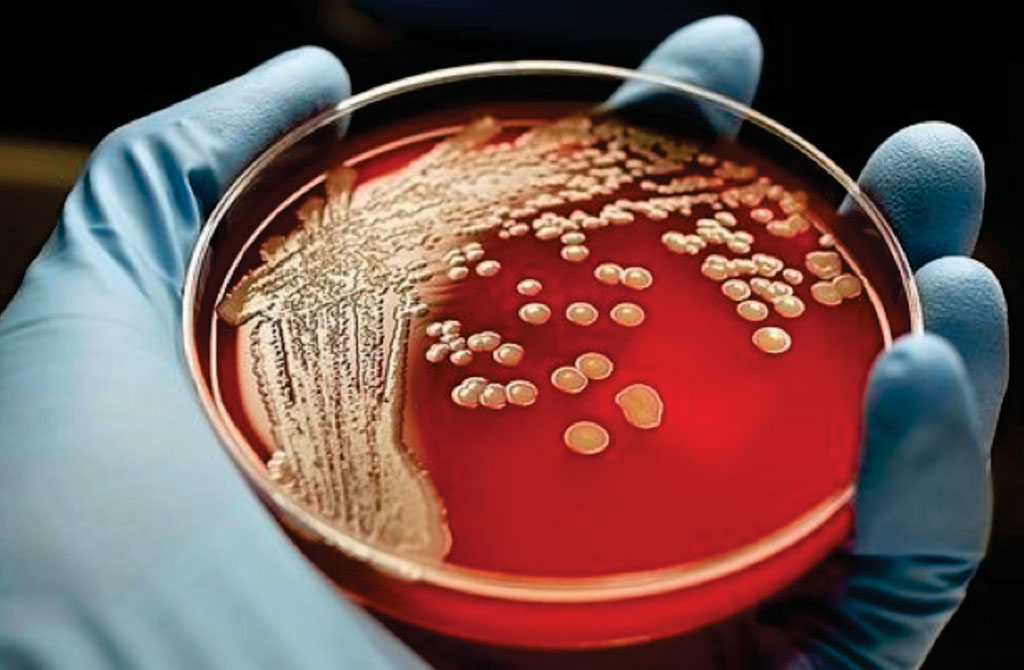
Image: Bacterial colonies of Staphylococcus aureus growing on horse blood agar (Photo courtesy of OMICS International).
A collaboration of scientists from the Centers for Disease Control and Prevention (Atlanta, GA USA) and the Georgia Institute of Technology (Atlanta, GA, USA) applied shotgun metagenomics to stool samples collected from two geographically isolated foodborne outbreaks in Alabama and Colorado, where the etiologic agents were identified as distinct strains of Salmonella enterica serovar Heidelberg by culture-dependent methods. The metagenomics data provided specific information about the bacterial phenotype involved and identified a secondary Staphylococcus aureus pathogen present in two of the samples tested. Knowing the specific phenotype can help in pinpointing the origins of an outbreak, while information about the secondary infection may help explain related factors such as the severity of the infection.
The scientists were also able to rule out one species, Escherichia coli (or E. coli), because the variant present was not of a virulent type. Variants of these bacteria are present naturally in the gut microbiome (called "commensal E. coli") while other variants are notorious enteric pathogens. Metagenomics showed the abundant E. coli population in the outbreak samples was probably commensal, and its growth may have been accelerated when conditions became more favorable during the Salmonella infection. In the two cases evaluated, scientists were able to determine that although the symptoms were similar, the outbreaks were caused by different variants of Salmonella and therefore were probably not connected.
Andrew D. Huang, PhD, a microbiologist/ bioinformatician and lead author of the study said, “Currently, the most advanced DNA fingerprinting method, whole genome sequencing, requires first pulling out, or isolating in a pure culture, the bacteria that made a person sick to generate a fingerprint. Metagenomics differs from whole genome sequencing because it could allow us to sequence the entire DNA in a patient's sample. It could allow us to skip the isolation steps and go directly from a stool sample to a highly detailed DNA fingerprint of the bacteria that made you sick. This method saves time and provides more detail that could be helpful for diagnosing a patient and identifying an outbreak.” The study was published on November 23, 2016, in the journal Applied and Environmental Microbiology.
Related Links:
Centers for Disease Control and Prevention
Georgia Institute of Technology