Multimodal AI Tool Predicts Genetic Alterations to Guide Breast Cancer Treatment
Posted on 25 Apr 2026
PIK3CA mutations are key biomarkers for selecting phosphoinositide 3-kinase (PI3K)–targeted therapies in breast cancer, yet access to molecular testing can be inconsistent and costly. Conventional polymerase chain reaction (PCR) and next-generation sequencing (NGS) require infrastructure that is not universally available. Single‑modality image-based predictors have also shown limited performance. A new study shows a cost-effective solution using an multimodal artificial intelligence (AI) framework that can infer PIK3CA mutation status from digital pathology images.
Hebei Medical University Fourth Hospital developed the Multimodal PIK3CA Model (MPM) to provide a cost‑effective, scalable, and reliable alternative to sequencing. The approach integrates whole‑slide imaging (WSI) with routinely captured clinical variables to produce a single prediction of mutation status. The framework was designed to address gaps left by models that rely on images or clinical data alone.
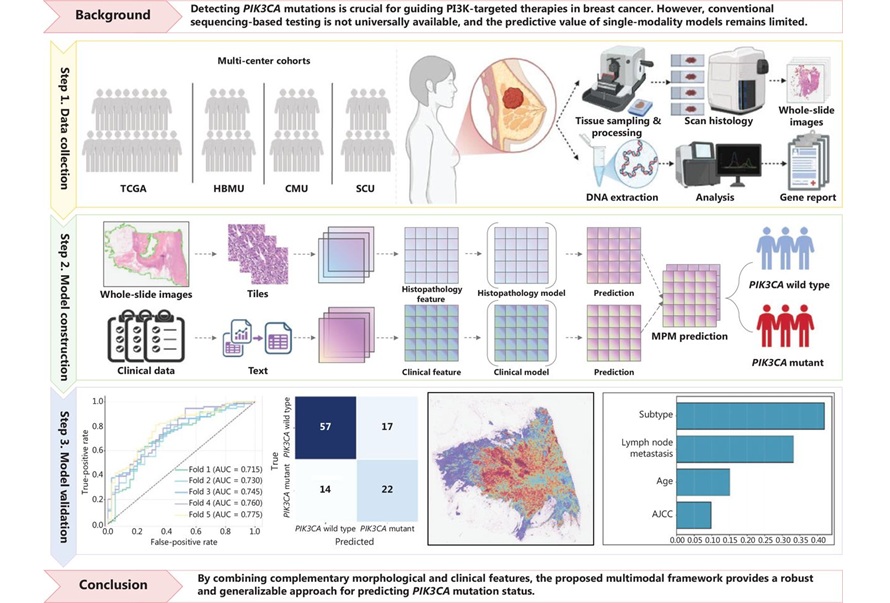
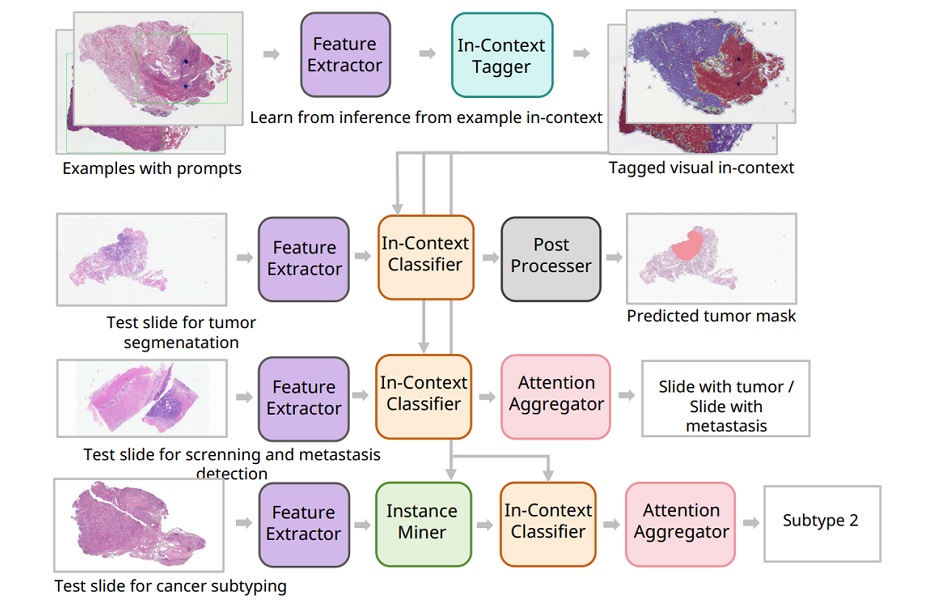
Within MPM, a histopathology model processes high‑resolution WSI using a transformer‑based pretrained encoder (H‑optimus‑0) and a clustering‑constrained attention multiple instance learning classifier (CLAM‑SB) to detect morphological correlates of PIK3CA mutations. A complementary clinical model based on XGBoost analyzes structured factors including age, molecular subtype, and lymph node status. The two models generate independent probabilities that are combined through decision‑level late fusion to yield the final classification.
The study drew on The Cancer Genome Atlas (TCGA) and three external clinical cohorts to test generalizability across institutions. In internal testing, the model achieved an area under the curve (AUC) of 0.745. External validation showed stable performance with AUCs ranging from 0.695 to 0.680, indicating robustness across diverse datasets.
According to the study, incorporating clinical variables improved predictive accuracy compared with single‑modality models. The authors describe the system as a practical option where conventional PCR or NGS is difficult to deploy, with potential utility for clinical decision support in varied settings. The work was published in Cancer Biology & Medicine on February 23, 2026.
The study notes that the framework demonstrated strong generalization and could be used in routine practice to help guide PI3K‑targeted treatment decisions. Future research may focus on refining the model for other mutations and additional cancer types.
“This multimodal AI framework represents a significant advancement in computational pathology. By integrating complementary clinical and morphological data, our model not only enhances the prediction of PIK3CA mutations but also offers a scalable, cost-effective solution for clinical practice. With its strong generalization across diverse cohorts, it has the potential to improve personalized treatment decisions for breast cancer patients, bridging the gap between advanced molecular testing and routine clinical workflows," said Dr. Yueping Liu, the lead corresponding author of the study.




.jpg)