New Gene Signature Reveals Underdiagnosed Lung Cancer Subtype
Posted on 13 Apr 2026
Combined small-cell lung cancer (cSCLC) is a rare lung malignancy in which tumors display both small-cell and non-small-cell features. These mixed tumors are commonly managed as standard small-cell disease even though patient outcomes are often worse, underscoring diagnostic and therapeutic challenges. Detection is further complicated when biopsy samples capture only a single tumor region, potentially missing the mixed nature of the disease. A new study sheds light on how these cancers arise and introduces a gene-based tool that may improve their identification.
At the Institute for Systems Biology (Seattle, WA, USA), investigators and collaborators developed a four-gene diagnostic tool, cSCLC Detector, to more accurately identify cSCLC. The tool emerged from a multi-institution effort co-led by the Institute for Systems Biology and focuses on distinguishing tumors that contain both small-cell and non-small-cell characteristics. In datasets of patients previously diagnosed with standard small-cell lung cancer (SCLC), the detector identified a substantially higher proportion of cases with combined features, indicating that cSCLC may be underdiagnosed.
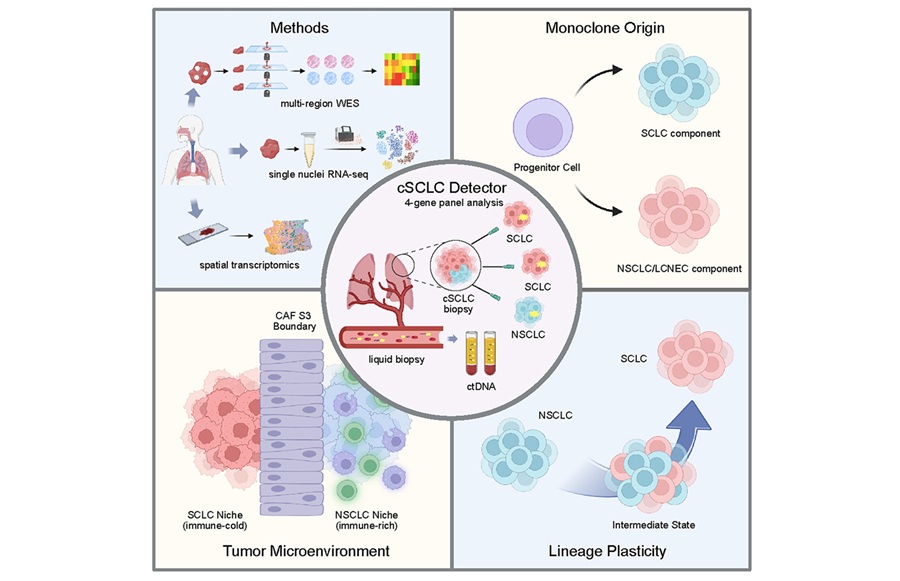
The detector is built on the observation that the small-cell and non-small-cell components of cSCLC share early trunk mutations and originate from the same ancestral tumor. Because of that shared genetic lineage, biopsy material that samples only a small-cell–appearing region can still carry genetic clues to a mixed identity. By leveraging a four-gene signature, the approach is designed to surface that hidden signal in routine analyses.
Using spatial and single-cell genomic approaches together with multi-region sequencing, the team found that cSCLC does not arise from two separate cancers. Instead, tumors originate from a single clone and can evolve over time, shifting between small-cell–like and non-small-cell–like states. Many cells occupy hybrid states in which they exhibit features of multiple cancer types simultaneously, with about one-third of analyzed small-cell–like cells showing these mixed identities.
In addition, the researchers found that different regions within the same tumor create distinct microenvironments. Some areas were rich in immune cells, while others were largely immune-excluded. Dense bands of fibroblasts—connective-tissue cells that tumors can co-opt to build protective barriers—often separated these regions. Those fibroblast-rich boundaries may help wall off parts of the tumor from immune attack.
The study is published in Cell Reports Medicine on April 10, 2026. Collaborating institutions include the Institute for Systems Biology and Fudan University. The findings emphasize the need to characterize how cancer cells change state and interact with their microenvironment, not solely their mutations.
“We found that these tumors are not simply mixtures of different cancer types. They are dynamic systems, with cancer cells actively changing their identity. That flexibility may help explain why they are so difficult to treat,” said Wei Wei, Ph.D., associate professor at ISB and co-corresponding author of the study.
“By combining spatial genomics, single-cell analysis, and multi-region sequencing, we were able to trace how these tumors evolve across both space and time. This approach allowed us to capture transitional cell states that are not visible using conventional methods,” said Qihui Shi, Ph.D., co-corresponding author at Fudan University.
Related Links
Institute for Systems Biology