Breakthrough Mass Spectrometry Design Could Enable Ultra-Low Abundance Detection
Posted on 31 Mar 2026
Mass spectrometry is central to identifying and quantifying molecules in complex biological samples, but conventional instruments typically analyze ions sequentially, which can limit detection of rare species. Researchers now describe a prototype that parallelizes ion handling to improve sensitivity for low-abundance targets in challenging matrices. The findings show enhanced dynamic range and signal-to-noise, potentially enabling detection of molecules that were previously undetectable with standard ion traps.
Most current systems examine one or a few ion populations at a time, a limitation that can slow analyses, raise costs, and obscure scarce but informative molecules in single-cell proteomics and metabolomics. Because proteins and metabolites cannot be amplified like nucleic acids, faint signals are easily overwhelmed by background. Improving sensitivity and throughput is therefore a recurring bottleneck for comprehensive molecular readouts.
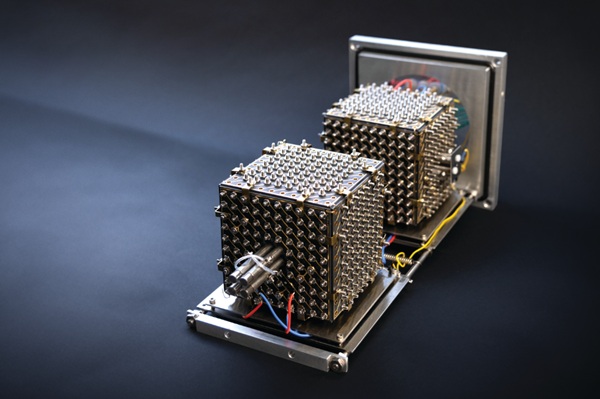

Investigators at Rockefeller University (New York, NY, USA) developed a prototype called MultiQ‑IT that cools, traps, filters, and redirects more than a billion ions simultaneously. Built to replace the core ion‑trapping stage, the cube‑shaped device is lined with hundreds of electrically controlled openings. Ions are slowed by collisions with residual gas, move stochastically within the chamber, and can be held or routed so that many ion populations are manipulated at once rather than one by one.
The team scaled the design from six openings to more than 1,000 and showed that a single incoming ion stream can be split into multiple parallel streams for simultaneous analysis. In a 486‑port configuration, MultiQ‑IT stored up to ten billion charges—about a thousand times the capacity of conventional ion traps. Applying a small voltage barrier across the exits allowed singly charged background ions to escape while retaining multiply charged, information‑rich ions, improving signal‑to‑noise by as much as 100‑fold and revealing previously undetectable proteins.
In a 1,134‑port design, just 39 open ports achieved half‑maximum depletion efficiency, echoing parallel transport seen in biological pores. The increased sensitivity could, for example, improve detection of low‑abundance crosslinked peptides used to map large protein complexes. The findings outline a physical blueprint for faster, more sensitive instruments and point toward routine, high‑throughput single‑cell proteomics. The work appears in Science Advances.
“It was the ability to run so many chemical reactions in parallel, which took genome sequencing from a billion-dollar effort to something that costs around $100. The same thing happened in computing with GPUs. And that’s what we’re trying to do with mass spectrometry,” said Brian T. Chait, the Camille and Henry Dreyfus Professor at The Rockefeller University.
Related Links
The Rockefeller University




.jpg)