New AI Tool Enables Rapid Treatment Selection in Pediatric Leukemia
Posted on 31 Mar 2026
Children with T-cell acute lymphoblastic leukemia face an aggressive disease that remains difficult to treat. Although remission rates have improved, many survivors experience long-term effects from intensive chemotherapy. Clinicians need faster ways to pinpoint which targeted therapies are likely to work for each patient to avoid ineffective regimens. Researchers now report an artificial intelligence-enabled microfluidic platform that predicts treatment response within hours.
At the University of Utah’s Huntsman Cancer Institute (Salt Lake City, UT, USA), scientists developed μPharma, an artificial intelligence (AI)-driven “lab-on-a-chip” system designed to rapidly forecast drug sensitivity for pediatric T-cell acute lymphoblastic leukemia (T-ALL). The tool, which is not yet used in clinical settings, produces results in under four hours, offering a potential pathway to same-day decision-making. According to the team, faster readouts could help reduce unnecessary treatments and side effects by identifying therapies to which a patient’s cancer is most responsive.
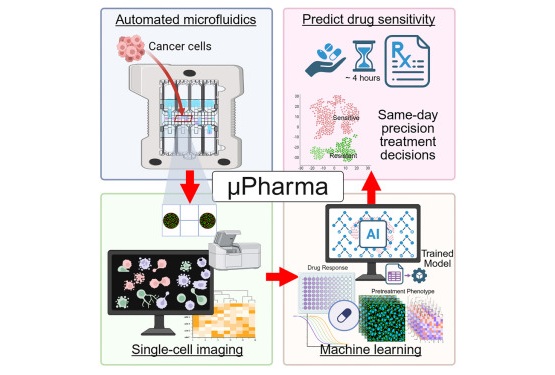
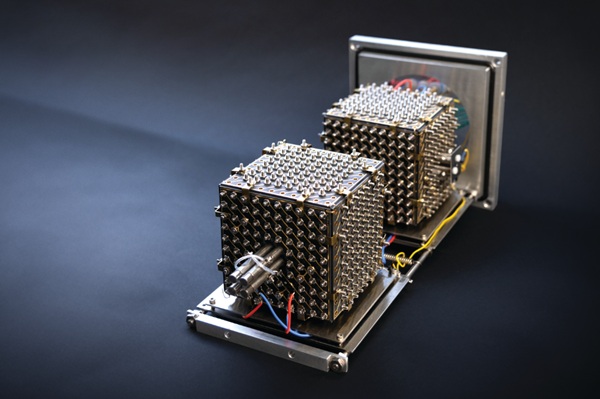
μPharma profiles drug response without directly exposing a patient’s cells to drugs. The device uses digital microfluidics to move droplets across the chip and automate liquid-handling steps, reducing cell and reagent requirements while minimizing human error. Within the cartridge, cancer cells are held between two closely spaced plates; electric currents shuttle reagents to and from the cells, revealing intracellular molecules linked to drug susceptibility. A machine learning model then analyzes molecular signals, their subcellular localization, and cell morphology to predict effective agents.
In a study published in Med on March 13, 2026, investigators reported that μPharma accurately predicted responses to two targeted therapies under investigation for T-ALL—dasatinib and venetoclax—and uncovered a previously unrecognized association between drug response and a key molecular marker in T-ALL. The platform evaluates heterogeneity at the single-cell level, enabling detection of subpopulations with differing susceptibilities that could drive relapse if left untreated. The project involved collaborators at St. Jude Children’s Research Hospital and the University of Pennsylvania.
“Innovation in treatment selection is a pressing need within pediatric malignancies. Personalized treatment selection accomplished in ‘real-time’ will be part of the future of cancer therapeutics, and μPharma represents an encouraging step in that direction,” said Luke Maese, DO, pediatric oncologist at Huntsman Cancer Institute and associate professor of pediatrics at the University of Utah.
“If we can rapidly and accurately monitor the sensitivity of cancer cells and tailor treatment appropriately, we believe it can significantly improve outcomes. The next step is validation of this technology using primary leukemia cells in a realistic clinical environment,” said Alphonsus Ng, Ph.D., assistant professor of biomedical engineering at the University of Utah and co-leader of the DigiPharma laboratory.
Related Links
Huntsman Cancer Institute




.jpg)