Proteomics-Based Diagnostic Test Predicts Risk of Dying from Staphylococcus aureus Bacteremia
By LabMedica International staff writers
Posted on 14 Sep 2020
A diagnostic approach that integrates proteomic and metabolomic techniques enabled the identification of biomarkers that could predict which patients had the highest risk of dying from Staphylococcus aureus bacteremia.Posted on 14 Sep 2020
Staphylococcus aureus bacteremia (SaB) causes significant disease in humans, carrying mortality rates of nearly 25%. The ability to rapidly predict SaB patient responses and guide personalized treatment regimens could reduce mortality.
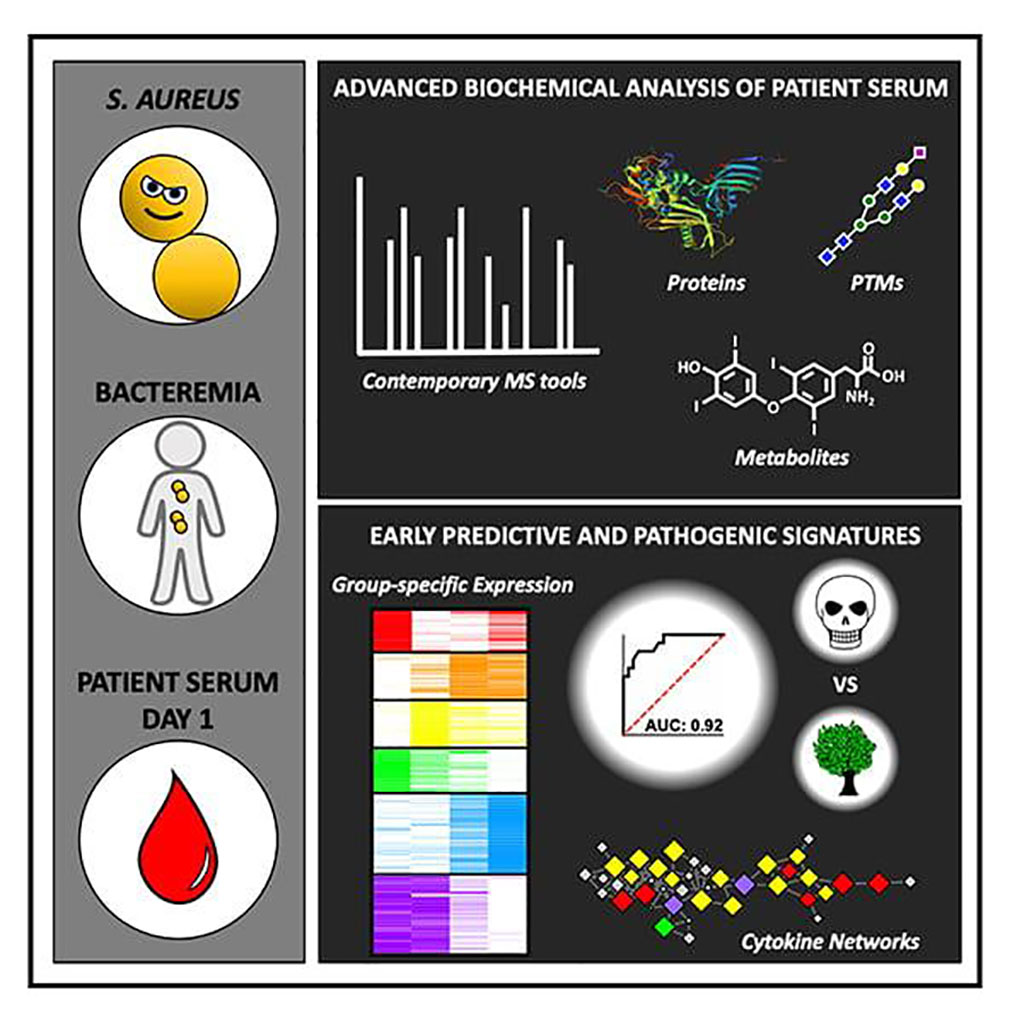
Image: Using advanced biochemical analysis of patient serum, researchers found signatures that help predict which patients are at highest risk of dying from S. aureus bacteremia, a blood infection (Photo courtesy of University of California, San Diego)
In order to develop this predictive capability, investigators at the University of California, San Diego (USA) utilized mass spectrometry (MS) to analyze more than 10,000 proteins and metabolites present in more than 200 serum samples collected from the blood of patients with SaB.
Results revealed that by integrating proteomic and metabolomic techniques, the investigators could identify more than 10,000 features from serum samples collected upon clinical presentation. The MS results demonstrated a specific pattern of proteins with and without post-translational modifications that differed in the serum of patients who ultimately died of SaB compared to those who did not. The biomarkers most highly associated with death included lower levels of glycosylated fetuin A, unmodified fetuin B, and thyroxine.
With the area under the curve (AUC) higher than 0.95, the biomarkers identified in this study greatly exceeded the predictive capabilities of those previously reported, particularly when used in combination.
"This finding is a leap forward toward a point-of-care predictive tool for bacteremia risk," said senior author Dr. David Gonzalez, assistant professor of medicine at the University of California, San Diego. "It also opens up lots of new basic biological questions about how our immune systems respond to infections. If I wanted to learn all about you, I would just talk to you directly, not your second cousin. Same thing here - we can gain new and important information by directly "asking" the proteins, rather than their genes, and mass spectrometry is currently the best way to do that in an unbiased manner."
The study was published in the September 3, 2020, online edition of the journal Cell.
Related Links:
University of California, San Diego









 (3) (1).png)