Magnetic Nanoparticle Tool Developed to Detect MicroRNA
By LabMedica International staff writers
Posted on 12 Sep 2018
A magnetic nanoparticle-based technique has been developed to directly detect microRNA (miRNA) in unprocessed whole blood samples in about thirty minutes. Currently scientists use quantitative reverse-transcription polymerase chain reaction (qRT-PCR) to profile miRNA due to its usually high specificity and sensitivity.Posted on 12 Sep 2018
Although qRT-PCR produces reliable results, scientists have noted that it does not detect miRNA directly in whole blood. Instead, scientists perform a lengthy process where they must isolate and purify RNA, and then synthesize complementary DNA (cDNA) to measure miRNA expression. The first human disease known to be associated with miRNA deregulation was chronic lymphocytic leukemia.
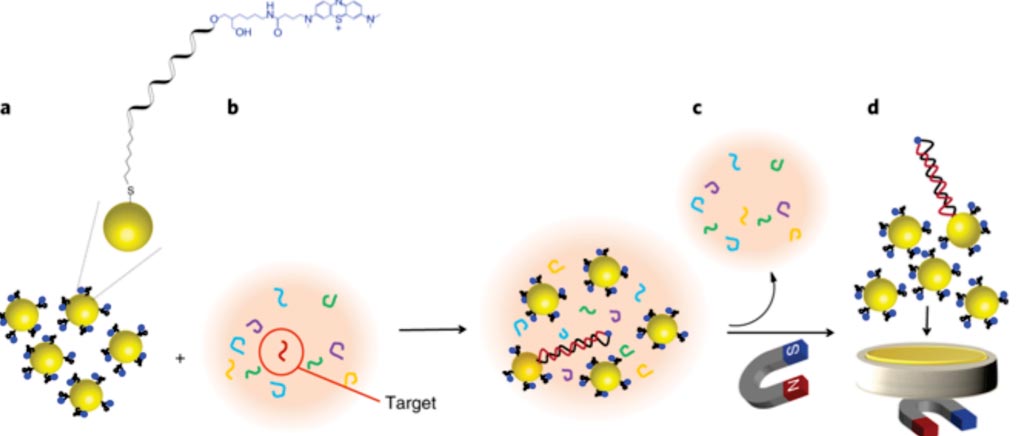
Image: Nucleic acid hybridization on an electrically reconfigurable network of gold-coated magnetic nanoparticles enables microRNA detection in blood (Photo courtesy of University of New South Wales).
Scientists at the University of New South Wales (Sydney, Australia) modified the surface of the gold-coated magnetic nanoparticles (Au@MNPs) with a probe DNA sequence complementary to miR-21 and marked it with a blue redox label. The team then added excess network of modified Au@MNPs by probe DNA (DNA-Au@NMPs) to the analyte solution. After 30 minutes, the team magnetically separated the Au@MNPs from the solution and washed away unhybridized sequences. They then collected the Au@MNPs on the surface of a gold microelectrode using a magnet. The team removed the magnet and applied 10 cycles of square-wave voltammetry between 200 and 500 megaVolts, with a pulse amplitude of 20 megavolts and frequency of 2 hertz. The team then used stable peak currents obtained before and after hybridization to measure the amount of analyte.
To examine the electrochemical tool's limit of detection and range of its sensor, the team prepared different concentrations of miR-21 in either phosphate buffered saline (PBS), undiluted human serum, or 50% whole human blood. The group measured a change in the current after 30 minutes of exposure to the miR-21 solutions within the concentration range of 10 attomolar to 1 nanomolar. In order to validate the electrochemical tool, the team used it to analyze miR-21 levels in a pool of other RNA sequences from the total RNA extracted from human lung cancer cells (A459) and exosomes. After treating the samples with an miR-21 inhibitor, the team still detected levels of miR-21 in exosomes eight times lower than the factor of 0.4 in their parental cells.
Using the DNA-Au@MNP-based electrochemical sensor, the team found that the concentration of miR-21 molecules in rapidly growing cancers was higher than those in the "no-control" group. The group also noted that the electrochemical assay had less variability than qRT-PCR. John Gooding, D. Phil, a chemistry professor and senior author of the study, said, “This tool would be especially useful for low levels of certain proteins and molecules, as we could potentially not only diagnose a patient's condition, but also look at treatment efficacy or the potential relapse of a condition like cancer. The study was published on August 27, 2018, in the journal Nature Nanotechnology.
Related Links:
University of New South Wales