Novel Method Reclaims Resolution of Single-Cell RNA-Seq
By LabMedica International staff writers
Posted on 28 Oct 2020
Single-cell RNA sequencing (scRNA-seq) is a powerful tool to characterize cells. Current scRNA-seq platforms, despite offering high throughput, are inefficient and provide low resolution among distinct cell states and molecular features. Posted on 28 Oct 2020
Most high-throughput scRNA-seq methods rely on barcoding of cellular components to recover single-cell transcriptomes for thousands of cells at once. This is achieved by isolating uniquely barcoded poly-dT oligonucleotides that can capture and tag cellular messenger RNA (mRNA) during reverse transcription. In a second step, an additional oligonucleotide priming site is added to newly synthesized complementary DNA (cDNA) to enable polymerase chain reaction (PCR)-based amplification.
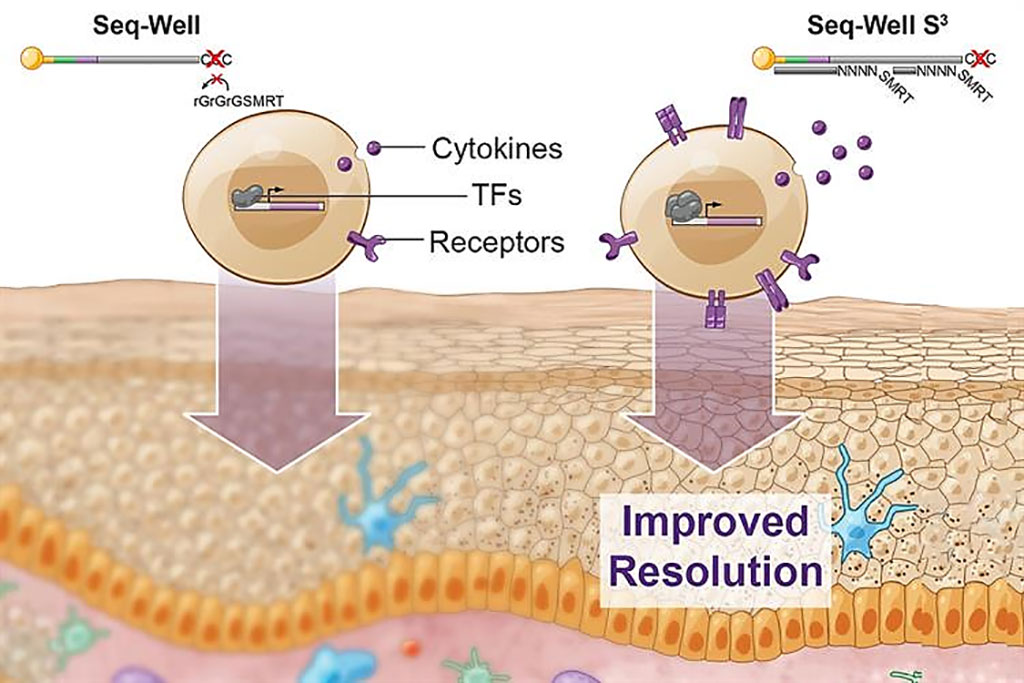
Image: Scientists have greatly boosted the amount of information that can be obtained using Seq-Well S3, a technique for rapidly sequencing RNA from single cells (Photo courtesy of MIT).
Medical Biochemists at the Massachusetts Institute of Technology (Cambridge, MA, USA) and their associates developed Seq-Well S3 ("Second-Strand Synthesis") as a massively parallel scRNA-seq protocol that uses a randomly primed second-strand synthesis to recover cDNA molecules to facilitate template-switching. This generates double-stranded cDNA that is labeled on one end with the SMART sequence and its reverse complement on the other, making it more accessible for PCR enzymes to amplify the molecules.
To perform the study skin biopsies were obtained from a total of 16 patients at the University of California, Los Angeles and University of Southern California Hansen’s Clinic, while an additional three samples were obtained from the University of Michigan. The team utilized Seq-Well, a massively parallel, low-input scRNA-seq platform for clinical samples, to capture the transcriptome of single cells. The team performed Templated Second-Strand Synthesis, PCR Amplification, Optimization of Second-Strand Synthesis, CD4+ T Cell comparisons of 10x Genomics, Pleasanton, CA, USA), Seq-Well S3, and Smart-Seq2, DNA Sequencing and Alignment of peripheral blood mononuclear cells (PBMC) optimization samples, and tissue immunofluorescence staining.
In total, the scientists processed 19 skin biopsies and retained over 38,000 high-quality single-cell transcriptomes using Seq-Well S3. They were able to recover 15 primary cell types. To further define biological features, the team used the method to examine subpopulations of T cells, myeloid cells, endothelial cells, dermal fibroblasts, and keratinocytes in each inflammatory condition. The team found, for example, regulatory T cells, dysfunctional NR4A1-expressing T cells, and senescent SESN3+ T cells were over-represented, potentially reflecting T-cell dysfunction in psoriasis pathology.
The team also distinguished patterns associated with multiple diseases by looking across different inflammatory skin conditions, revealing common and unique features. For instance, they found that a group of natural killer cells, γΔ T cells, and a sub-cluster of immature cytotoxic T cells are derived from leprosy and granuloma annulare, indicating common T-cell programming in both forms of inflammation.
Alex Shalek, PhD, associate professor of chemistry at MIT and a senior author of the study, said, “"It's become clear that these technologies have transformative potential for understanding complex biological systems. If we look across a range of different datasets, we can really understand the landscape of health and disease, and that can give us information as to what therapeutic strategies we might employ.” The study was published on October 13, 2020 in the journal Immunity.
Related Links:
Massachusetts Institute of Technology
10x Genomics