NGS Identifies Genomic Characteristics of Pediatric Leukemia
By LabMedica International staff writers
Posted on 19 Mar 2020
Pediatric leukemias have a diverse genomic landscape associated with complex structural variants, including gene fusions, insertions and deletions, and single nucleotide variants. Routine karyotype and fluorescence in situ hybridization (FISH) techniques lack sensitivity for smaller genomic alternations.Posted on 19 Mar 2020
Minimal residual disease (MRD; more aptly named Measurable Residual Disease) is the detection of residual leukemia following therapy, most commonly by flow cytometry. MRD, as measured by multiparameter flow cytometry is perhaps one of the most important predictors of outcome in children with acute myelogenous leukemia.
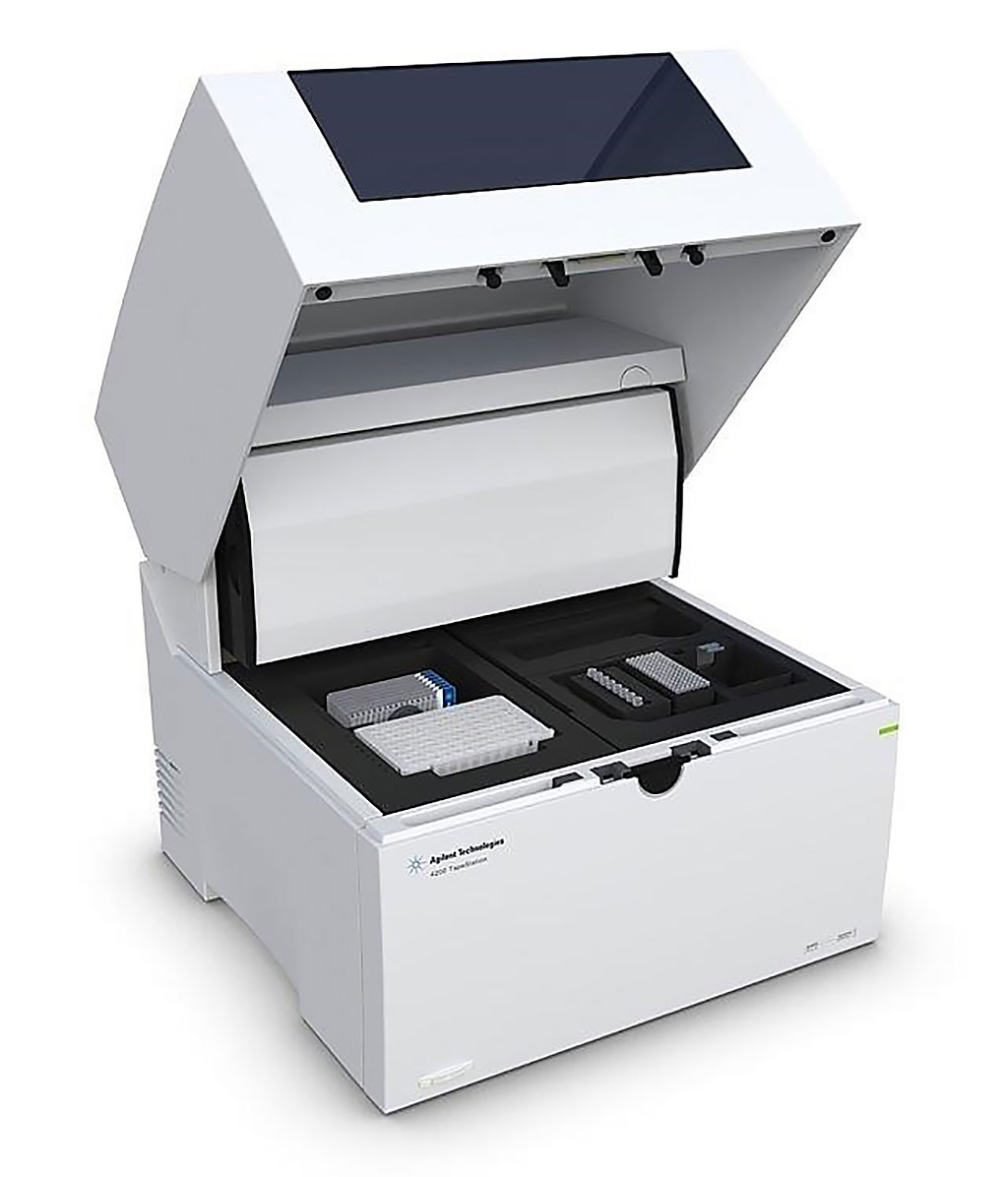

Image: The Agilent 4200 TapeStation system is an established automated electrophoresis tool for DNA and RNA sample quality control (Photo courtesy of Agilent Technologies).
Hematologists at the Nemours/Alfred I. duPont Hospital for Children (Wilmington, DE, USA) and their associates collected 32 primary bone marrow samples from pediatric leukemia and five adult leukemia subjects, cell line MV4–11, and an umbilical cord sample. Patient samples were collected at diagnosis, end of the first treatment, and relapse. The Nemours samples consisted of six acute myelogenous leukemia (AML) subjects, one acute promyelocytic leukemia (APL) subject, 17 preB-cell acute lymphoblastic leukemia (ALL) subjects and three T-cell ALL subjects.
Nucleic acid was extracted from each sample. DNA was extracted using the DNeasy Blood & Tissue Kit (Qiagen, Germantown, MD, USA). Nucleic acid quantity and quality was then assessed using the TapeStation 4200 (Agilent Technologies, Santa Clara, CA, USA). To optimize detection of structural and copy number variants in DNA and RNA in genes more closely affiliated with pediatric leukemia, the team prepared DNA- error-corrected sequencing (ECS) libraries using a customized VariantPlex kit (ArcherDx, Boulder, CO, USA) or a human cancer transcriptome assay.
The scientists reported that similar to flow cytometry for ALL MRD, the limit of detection (LOD) for point mutations by their sequencing strategies was ≥0.001. For DNA structural variants, FLT3 internal tandem duplication (ITD) positive cell line and patient samples showed a LOD of ≥0.001 in addition to previously unknown copy number losses in leukemia genes. ECS in RNA identified multiple novel gene fusions, including a SPANT-ABL gene fusion in an ALL patient, which could have been used to alter therapy.
Collectively, ECS for RNA demonstrated a quantitative and complex landscape of RNA molecules with 12% of the molecules representing gene fusions, 12% exon duplications, 8% exon deletions, and 68% with retained introns. Droplet digital polymerase chain reaction validation of ECS-RNA confirmed results to single mRNA molecule quantities.
The authors concluded that collectively, the assays enabled a highly sensitive, comprehensive, and simultaneous analysis of various clonal leukemic mutations, which can be tracked across disease states (diagnosis, end of induction, and relapse) with a high degree of sensitivity. The study was published on March 4, 2020 in the journal BMC Medical Genomics.
Related Links:
Nemours/Alfred I. duPont Hospital for Children
Qiagen
Agilent Technologies
ArcherDx