Residual Tissue Repositories Suitable for MS-Based Proteomics
By LabMedica International staff writers
Posted on 12 Sep 2018
Mass spectrometry-based proteomics has become a powerful tool for the identification and quantification of proteins from a wide variety of biological specimens.Posted on 12 Sep 2018
The majority of studies utilizing tissue samples have been carried out on prospectively collected fresh frozen or optimal cutting temperature (OCT) embedded specimens. However, such specimens are often difficult to obtain, in limited in supply, and clinical information and outcomes on patients are inherently delayed as compared to banked samples.
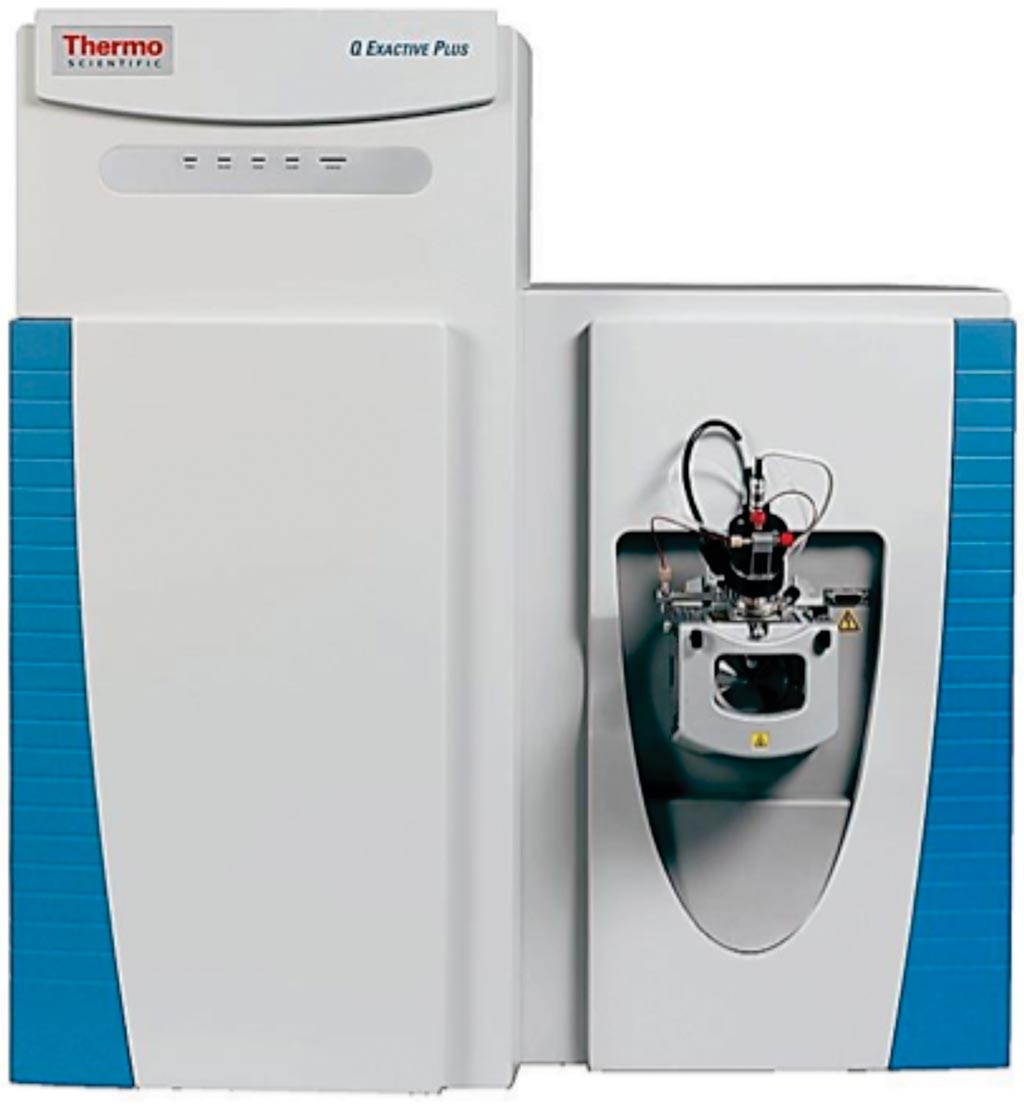

Image: The Q Exactive hybrid quadrupole-orbitrap mass spectrometer (Photo courtesy of Thermo Fisher Scientific).
Scientists from the Pacific Northwest National Laboratory (Richland, WA, USA) and their colleagues analyzed 60 patient samples taken from the NCI's Surveillance, Epidemiology, and End Results (SEER) residual tissue repositories, which contain samples from more than 100,000 cancer patients, along with detailed demographic information, data on tumor characteristics, treatment, survival, and cause of death. The 60 samples ranged in their time of storage from seven to 32 years.
The team used 10-plex tandem mass tag (TMT) labeling and divided each sample into six fractions, each of which they then ran on a 100-minute nanoLC gradient upfront of analysis on a Q-Exactive Plus instrument. For phosphopeptide analysis, they used Immobilized Metal Affinity Chromatography (IMAC) enrichment. They found that all 60 samples provided sufficient material for proteome-wide protein expression analysis and 18 of the 60 samples provided enough material for phosphopeptide work.
The investigators identified and quantified a total of 8,582 proteins and 8,073 phosphopeptides across the SEER sample set, indicating that FFPE tissue is amenable to mass spec proteomics analysis. Protein identifications were reduced compared to the identifications possible in comparable optimal cutting temperature (OCT) compound-embedded specimens. Compared to OCT samples, peptide, protein, and phosphopeptide identifications were reduced by 50%, 20%, and 76%, respectively.
Karin D, Rodland, PhD, an expert in mass spectrometry and a senior author of the study, said, “There have been commercial kits available for 12 to 15 years for extracting proteins out of FFPE blocks, and on the face of it, the yields of protein from the FFPE blocks is not that bad. But with the mass spec technologies of 12 to 15 years ago, the rate of identification was very low. You just didn't get good [proteome] coverage out of FFPE blocks. And the assumption was that the formalin crosslinking was causing you to lose identifications. However, improvements in mass spec technology have provided instruments with higher sensitivity and better resolution that are capable of working with smaller amounts of sample.” The study was published on August 3, 2018, in the journal Clinical Proteomics.
Related Links:
Pacific Northwest National Laboratory