Fecal Microbes Used to Diagnose Liver Cirrhosis
|
By LabMedica International staff writers Posted on 09 Apr 2019 |
Image: The UltraClean PCR cleanup kit (Photo courtesy of MO BIO Laboratories).
Nonalcoholic fatty liver disease (NAFLD) is the most prevalent cause of chronic liver disease worldwide, and yet remains largely underdiagnosed even in individuals with advanced stage of the disease.
NAFLD-cirrhosis represents the most severe stage of the disease, carries a significant risk of hepatocellular carcinoma (HCC), and is consistently identified as the most important predictor of liver-related morbidity-mortality in NAFLD.
Medical scientists at the University of California San Diego (La Jolla, CA, USA) and their associates analyzed the microbial makeup of stool samples from 98 people known to have some form of NAFLD and 105 of their first-degree relatives, including some twins. The study included 26 probands with NAFLD-cirrhosis and 37 of their first-degree relatives. At the time of each visit, patients provided stool samples. These were collected and immediately stored in a −80 °C freezer.
DNA extraction and 16S rRNA amplicon sequencing were done and DNA was extracted using the Qiagen MagAttract PowerSoil DNA kit. Amplicon PCR was performed on the V4 region of the 16S rRNA gene using the primer pair 515 f to 806r with Golay error-correcting barcodes on the reverse primer. Amplicons were barcoded and pooled in equal concentrations for sequencing. The amplicon pool was purified with the MO BIO UltraClean PCR cleanup kit and sequenced on the MiSeq sequencing platform.
The team identified 27 unique bacterial features unique to the gut microbiomes from stools of people with NAFLD-cirrhosis. They were able to use this noninvasive stool test to pick out the people with known NAFLD-cirrhosis with 92% accuracy, but more importantly, the test allowed them to differentiate the first-degree relative with previously undiagnosed NAFLD-cirrhosis with 87% accuracy. The results were confirmed by magnetic resonance imaging (MRI).
At the genus level, both NAFLD-cirrhosis and NAFLD without advanced fibrosis group were enriched with Streptococcus, but only the NAFLD-cirrhosis group was enriched with Megasphaera. Species belonging to the family Enterobacteriaceae and the genera Streptococcus and Gallibacterium were the most enriched in NAFLD-cirrhosis, while Faecalibacterium prausnitzii and species belonging to the genus, Catenibacterium and the families Rikenellaceae, Mogibacterium, Peptostreptococcaceae were enriched in non-NAFLD controls.
Rohit Loomba, MD, a professor of medicine and senior author of the study, said, “If we are better able to diagnose NAFLD-related cirrhosis, we will be better at enrolling the right types of patients in clinical trials, and ultimately will be better equipped to prevent and treat it. This latest advance toward a noninvasive stool test for NAFLD-cirrhosis may also help pave the way for other microbiome-based diagnostics and therapeutics, and better enable us to provide personalized, or precision, medicine for a number of conditions.” The study was published on March 29, 2019, in the journal Nature Communications.
Related Links:
University of California San Diego
NAFLD-cirrhosis represents the most severe stage of the disease, carries a significant risk of hepatocellular carcinoma (HCC), and is consistently identified as the most important predictor of liver-related morbidity-mortality in NAFLD.
Medical scientists at the University of California San Diego (La Jolla, CA, USA) and their associates analyzed the microbial makeup of stool samples from 98 people known to have some form of NAFLD and 105 of their first-degree relatives, including some twins. The study included 26 probands with NAFLD-cirrhosis and 37 of their first-degree relatives. At the time of each visit, patients provided stool samples. These were collected and immediately stored in a −80 °C freezer.
DNA extraction and 16S rRNA amplicon sequencing were done and DNA was extracted using the Qiagen MagAttract PowerSoil DNA kit. Amplicon PCR was performed on the V4 region of the 16S rRNA gene using the primer pair 515 f to 806r with Golay error-correcting barcodes on the reverse primer. Amplicons were barcoded and pooled in equal concentrations for sequencing. The amplicon pool was purified with the MO BIO UltraClean PCR cleanup kit and sequenced on the MiSeq sequencing platform.
The team identified 27 unique bacterial features unique to the gut microbiomes from stools of people with NAFLD-cirrhosis. They were able to use this noninvasive stool test to pick out the people with known NAFLD-cirrhosis with 92% accuracy, but more importantly, the test allowed them to differentiate the first-degree relative with previously undiagnosed NAFLD-cirrhosis with 87% accuracy. The results were confirmed by magnetic resonance imaging (MRI).
At the genus level, both NAFLD-cirrhosis and NAFLD without advanced fibrosis group were enriched with Streptococcus, but only the NAFLD-cirrhosis group was enriched with Megasphaera. Species belonging to the family Enterobacteriaceae and the genera Streptococcus and Gallibacterium were the most enriched in NAFLD-cirrhosis, while Faecalibacterium prausnitzii and species belonging to the genus, Catenibacterium and the families Rikenellaceae, Mogibacterium, Peptostreptococcaceae were enriched in non-NAFLD controls.
Rohit Loomba, MD, a professor of medicine and senior author of the study, said, “If we are better able to diagnose NAFLD-related cirrhosis, we will be better at enrolling the right types of patients in clinical trials, and ultimately will be better equipped to prevent and treat it. This latest advance toward a noninvasive stool test for NAFLD-cirrhosis may also help pave the way for other microbiome-based diagnostics and therapeutics, and better enable us to provide personalized, or precision, medicine for a number of conditions.” The study was published on March 29, 2019, in the journal Nature Communications.
Related Links:
University of California San Diego
Latest Molecular Diagnostics News
- Blood Test Predicts Knee Osteoarthritis Eight Years Before Signs Appears On X-Rays
- Blood Test Accurately Predicts Lung Cancer Risk and Reduces Need for Scans
- Unique Autoantibody Signature to Help Diagnose Multiple Sclerosis Years before Symptom Onset
- Blood Test Could Detect HPV-Associated Cancers 10 Years before Clinical Diagnosis
- Low-Cost Point-Of-Care Diagnostic to Expand Access to STI Testing
- 18-Gene Urine Test for Prostate Cancer to Help Avoid Unnecessary Biopsies
- Urine-Based Test Detects Head and Neck Cancer
- Blood-Based Test Detects and Monitors Aggressive Small Cell Lung Cancer
- Blood-Based Machine Learning Assay Noninvasively Detects Ovarian Cancer
- Simple PCR Assay Accurately Differentiates Between Small Cell Lung Cancer Subtypes
- Revolutionary T-Cell Analysis Approach Enables Cancer Early Detection
- Single Genetic Test to Accelerate Diagnoses for Rare Developmental Disorders
- Upgraded Syndromic Testing Analyzer Enables Remote Test Results Access
- Respiratory and Throat Infection PCR Test Detects Multiple Pathogens with Overlapping Symptoms
- Blood Circulating Nucleic Acid Enrichment Technique Enables Non-Invasive Liver Cancer Diagnosis
- First FDA-Approved Molecular Test to Screen Blood Donors for Malaria Could Improve Patient Safety
Channels
Clinical Chemistry
view channel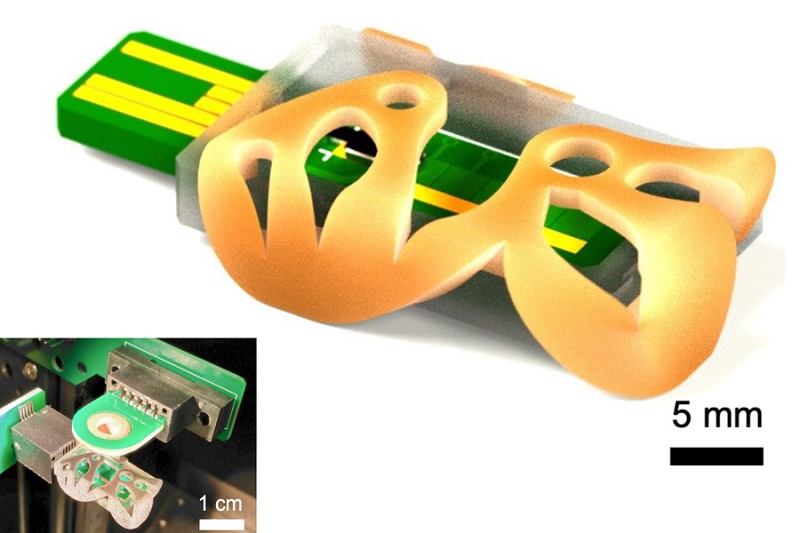
3D Printed Point-Of-Care Mass Spectrometer Outperforms State-Of-The-Art Models
Mass spectrometry is a precise technique for identifying the chemical components of a sample and has significant potential for monitoring chronic illness health states, such as measuring hormone levels... Read more.jpg)
POC Biomedical Test Spins Water Droplet Using Sound Waves for Cancer Detection
Exosomes, tiny cellular bioparticles carrying a specific set of proteins, lipids, and genetic materials, play a crucial role in cell communication and hold promise for non-invasive diagnostics.... Read more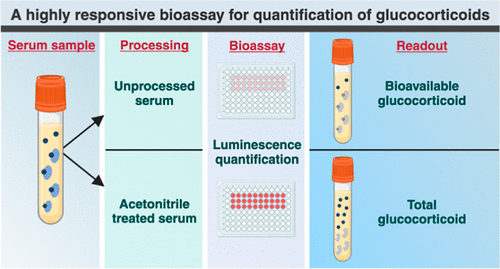
Highly Reliable Cell-Based Assay Enables Accurate Diagnosis of Endocrine Diseases
The conventional methods for measuring free cortisol, the body's stress hormone, from blood or saliva are quite demanding and require sample processing. The most common method, therefore, involves collecting... Read moreHematology
view channel
Next Generation Instrument Screens for Hemoglobin Disorders in Newborns
Hemoglobinopathies, the most widespread inherited conditions globally, affect about 7% of the population as carriers, with 2.7% of newborns being born with these conditions. The spectrum of clinical manifestations... Read more
First 4-in-1 Nucleic Acid Test for Arbovirus Screening to Reduce Risk of Transfusion-Transmitted Infections
Arboviruses represent an emerging global health threat, exacerbated by climate change and increased international travel that is facilitating their spread across new regions. Chikungunya, dengue, West... Read more
POC Finger-Prick Blood Test Determines Risk of Neutropenic Sepsis in Patients Undergoing Chemotherapy
Neutropenia, a decrease in neutrophils (a type of white blood cell crucial for fighting infections), is a frequent side effect of certain cancer treatments. This condition elevates the risk of infections,... Read more
First Affordable and Rapid Test for Beta Thalassemia Demonstrates 99% Diagnostic Accuracy
Hemoglobin disorders rank as some of the most prevalent monogenic diseases globally. Among various hemoglobin disorders, beta thalassemia, a hereditary blood disorder, affects about 1.5% of the world's... Read moreImmunology
view channel
Diagnostic Blood Test for Cellular Rejection after Organ Transplant Could Replace Surgical Biopsies
Transplanted organs constantly face the risk of being rejected by the recipient's immune system which differentiates self from non-self using T cells and B cells. T cells are commonly associated with acute... Read more
AI Tool Precisely Matches Cancer Drugs to Patients Using Information from Each Tumor Cell
Current strategies for matching cancer patients with specific treatments often depend on bulk sequencing of tumor DNA and RNA, which provides an average profile from all cells within a tumor sample.... Read more
Genetic Testing Combined With Personalized Drug Screening On Tumor Samples to Revolutionize Cancer Treatment
Cancer treatment typically adheres to a standard of care—established, statistically validated regimens that are effective for the majority of patients. However, the disease’s inherent variability means... Read moreMicrobiology
view channel
New CE-Marked Hepatitis Assays to Help Diagnose Infections Earlier
According to the World Health Organization (WHO), an estimated 354 million individuals globally are afflicted with chronic hepatitis B or C. These viruses are the leading causes of liver cirrhosis, liver... Read more
1 Hour, Direct-From-Blood Multiplex PCR Test Identifies 95% of Sepsis-Causing Pathogens
Sepsis contributes to one in every three hospital deaths in the US, and globally, septic shock carries a mortality rate of 30-40%. Diagnosing sepsis early is challenging due to its non-specific symptoms... Read morePathology
view channel
First of Its Kind Universal Tool to Revolutionize Sample Collection for Diagnostic Tests
The COVID pandemic has dramatically reshaped the perception of diagnostics. Post the pandemic, a groundbreaking device that combines sample collection and processing into a single, easy-to-use disposable... Read moreAI-Powered Digital Imaging System to Revolutionize Cancer Diagnosis
The process of biopsy is important for confirming the presence of cancer. In the conventional histopathology technique, tissue is excised, sliced, stained, mounted on slides, and examined under a microscope... Read more
New Mycobacterium Tuberculosis Panel to Support Real-Time Surveillance and Combat Antimicrobial Resistance
Tuberculosis (TB), the leading cause of death from an infectious disease globally, is a contagious bacterial infection that primarily spreads through the coughing of patients with active pulmonary TB.... Read moreTechnology
view channel
New Diagnostic System Achieves PCR Testing Accuracy
While PCR tests are the gold standard of accuracy for virology testing, they come with limitations such as complexity, the need for skilled lab operators, and longer result times. They also require complex... Read more
DNA Biosensor Enables Early Diagnosis of Cervical Cancer
Molybdenum disulfide (MoS2), recognized for its potential to form two-dimensional nanosheets like graphene, is a material that's increasingly catching the eye of the scientific community.... Read more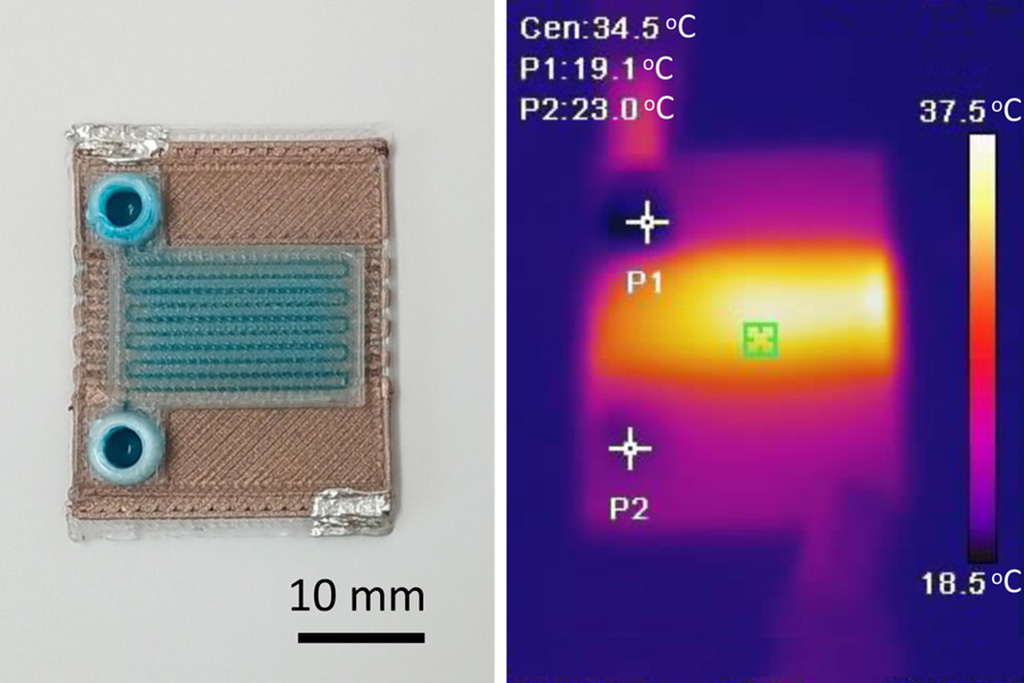
Self-Heating Microfluidic Devices Can Detect Diseases in Tiny Blood or Fluid Samples
Microfluidics, which are miniature devices that control the flow of liquids and facilitate chemical reactions, play a key role in disease detection from small samples of blood or other fluids.... Read more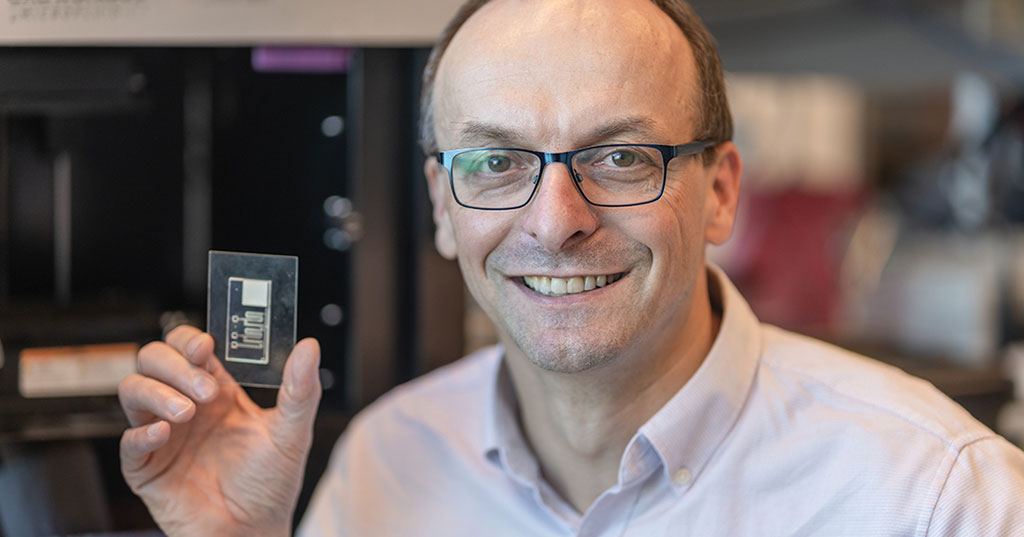
Breakthrough in Diagnostic Technology Could Make On-The-Spot Testing Widely Accessible
Home testing gained significant importance during the COVID-19 pandemic, yet the availability of rapid tests is limited, and most of them can only drive one liquid across the strip, leading to continued... Read moreIndustry
view channel
ECCMID Congress Name Changes to ESCMID Global
Over the last few years, the European Society of Clinical Microbiology and Infectious Diseases (ESCMID, Basel, Switzerland) has evolved remarkably. The society is now stronger and broader than ever before... Read more
Bosch and Randox Partner to Make Strategic Investment in Vivalytic Analysis Platform
Given the presence of so many diseases, determining whether a patient is presenting the symptoms of a simple cold, the flu, or something as severe as life-threatening meningitis is usually only possible... Read more
Siemens to Close Fast Track Diagnostics Business
Siemens Healthineers (Erlangen, Germany) has announced its intention to close its Fast Track Diagnostics unit, a small collection of polymerase chain reaction (PCR) testing products that is part of the... Read more











.jpg)
