Cancerous Tissue Analysis Introduced Using On-chip Technology
|
By LabMedica International staff writers Posted on 29 Jan 2018 |
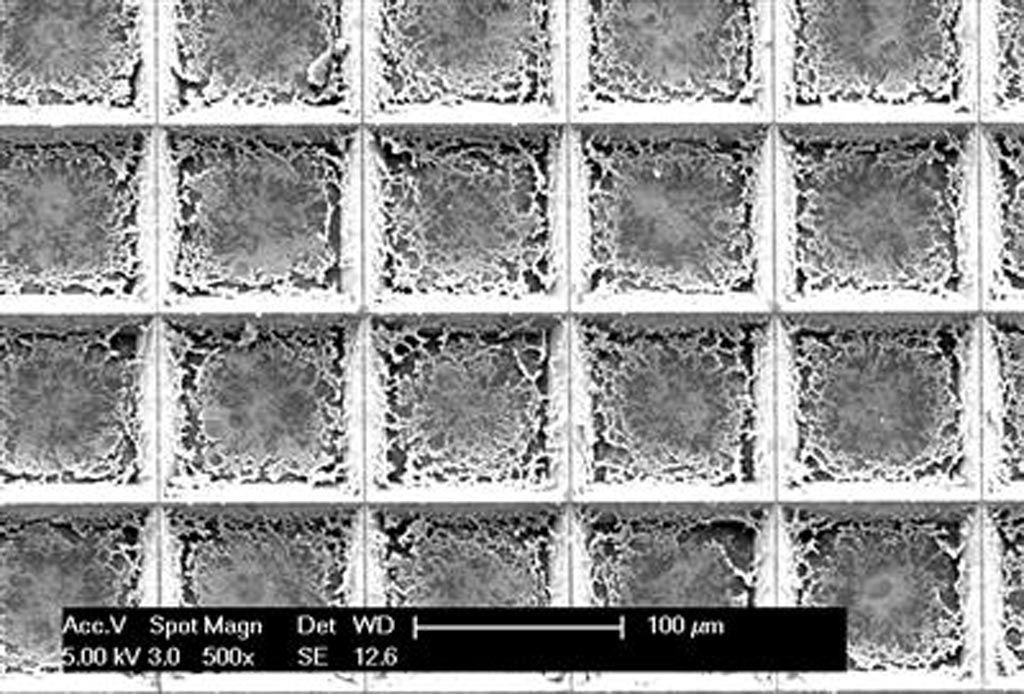

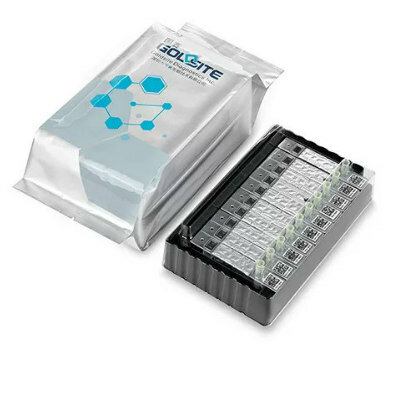
Image: The pixelated spatial gene expression tool can analyze an entire tissue sample and identify cancer cells in a process that takes less than two hours (Photo courtesy of the University of Illinois).
A novel gene expression analysis technique has been introduced that can accurately measure levels of RNA quickly and directly from a cancerous tissue sample while preserving the spatial information across the tissue.
The new technique performs on-chip picoliter real-time reverse transcriptase loop mediated isothermal amplification (RT-LAMP) reactions on a histological tissue section without any analyte purification while preserving the native spatial location of the nucleic acid molecules.
LAMP is an isothermal nucleic acid amplification technique, which in contrast to polymerase chain reaction (PCR) technology in which the reaction is carried out with a series of alternating temperature steps or cycles, is carried out at a constant temperature and does not require a thermal cycler. In LAMP, the target sequence is amplified at a constant temperature of 60–65 degrees Celsius using either two or three sets of primers and a polymerase with high strand displacement activity in addition to a replication activity. An additional pair of "loop primers" can further accelerate the reaction. Due to the specific nature of the action of these primers, the amount of DNA produced in LAMP is considerably higher than PCR based amplification.
The new technique developed through collaboration between the University of Illinois (Champaign-Urbana, USA) and the Mayo Clinic (Rochester, MN, USA) is conducted on a fingernail-size silicon chip that contains an array of more than 5,000 pyramid-shaped wells with razor-sharp edges. When a centimeter-sized cancer tissue sample is placed on the chip it is automatically cut up into hundreds or thousands of tiny pieces (in a two minute process called “tissue pixelation”). This step is followed by tissue fixation (10 minutes), permeabilization (30 minutes), loading of wells with amplification reagents (two minutes), and finally on-chip picoliter RT-LAMP reaction on a hot plate (45 minutes).
The investigators demonstrated this method by amplifying TOP2A messenger RNA (mRNA) in a prostate cancer xenograft with 100-micron spatial resolution and by visualizing the variation in threshold time of amplification across the tissue. The on-chip reaction was validated by mRNA fluorescence in situ hybridization (mFISH) from cells in the tissue section. The entire process, from tissue loading on microchip to results from RT-LAMP could be carried out in less than two hours.
"Our approach pixelates the entire tissue sample and can identify those very few cells that may be cancerous," said senior author Dr. Rashid Bashir, professor of bioengineering at the University of Illinois. "There is not any technique in existence that takes raw tissue to nucleic acid amplification while keeping the spatial information preserved."
Details of the new technique were published in the January 15, 2018, online edition of the journal Nature Communications.
Related Links:
University of Illinois
Mayo Clinic
The new technique performs on-chip picoliter real-time reverse transcriptase loop mediated isothermal amplification (RT-LAMP) reactions on a histological tissue section without any analyte purification while preserving the native spatial location of the nucleic acid molecules.
LAMP is an isothermal nucleic acid amplification technique, which in contrast to polymerase chain reaction (PCR) technology in which the reaction is carried out with a series of alternating temperature steps or cycles, is carried out at a constant temperature and does not require a thermal cycler. In LAMP, the target sequence is amplified at a constant temperature of 60–65 degrees Celsius using either two or three sets of primers and a polymerase with high strand displacement activity in addition to a replication activity. An additional pair of "loop primers" can further accelerate the reaction. Due to the specific nature of the action of these primers, the amount of DNA produced in LAMP is considerably higher than PCR based amplification.
The new technique developed through collaboration between the University of Illinois (Champaign-Urbana, USA) and the Mayo Clinic (Rochester, MN, USA) is conducted on a fingernail-size silicon chip that contains an array of more than 5,000 pyramid-shaped wells with razor-sharp edges. When a centimeter-sized cancer tissue sample is placed on the chip it is automatically cut up into hundreds or thousands of tiny pieces (in a two minute process called “tissue pixelation”). This step is followed by tissue fixation (10 minutes), permeabilization (30 minutes), loading of wells with amplification reagents (two minutes), and finally on-chip picoliter RT-LAMP reaction on a hot plate (45 minutes).
The investigators demonstrated this method by amplifying TOP2A messenger RNA (mRNA) in a prostate cancer xenograft with 100-micron spatial resolution and by visualizing the variation in threshold time of amplification across the tissue. The on-chip reaction was validated by mRNA fluorescence in situ hybridization (mFISH) from cells in the tissue section. The entire process, from tissue loading on microchip to results from RT-LAMP could be carried out in less than two hours.
"Our approach pixelates the entire tissue sample and can identify those very few cells that may be cancerous," said senior author Dr. Rashid Bashir, professor of bioengineering at the University of Illinois. "There is not any technique in existence that takes raw tissue to nucleic acid amplification while keeping the spatial information preserved."
Details of the new technique were published in the January 15, 2018, online edition of the journal Nature Communications.
Related Links:
University of Illinois
Mayo Clinic
Latest Molecular Diagnostics News
- Game-Changing Blood Test for Stroke Detection Could Bring Life-Saving Care to Patients
- Blood Proteins Could Warn of Cancer Seven Years before Diagnosis
- New DNA Origami Technique to Advance Disease Diagnosis
- Ultrasound-Aided Blood Testing Detects Cancer Biomarkers from Cells
- New Respiratory Syndromic Testing Panel Provides Fast and Accurate Results
- New Synthetic Biomarker Technology Differentiates Between Prior Zika and Dengue Infections
- Novel Biomarkers to Improve Diagnosis of Renal Cell Carcinoma Subtypes
- RNA-Powered Molecular Test to Help Combat Early-Age Onset Colorectal Cancer
- Advanced Blood Test to Spot Alzheimer's Before Progression to Dementia
- Multi-Omic Noninvasive Urine-Based DNA Test to Improve Bladder Cancer Detection
- First of Its Kind NGS Assay for Precise Detection of BCR::ABL1 Fusion Gene to Enable Personalized Leukemia Treatment
- Urine Test to Revolutionize Lyme Disease Testing
- Simple Blood Test Could Enable First Quantitative Assessments for Future Cerebrovascular Disease
- New Genetic Testing Procedure Combined With Ultrasound Detects High Cardiovascular Risk
- Blood Samples Enhance B-Cell Lymphoma Diagnostics and Prognosis
- Blood Test Predicts Knee Osteoarthritis Eight Years Before Signs Appears On X-Rays
Channels
Clinical Chemistry
view channel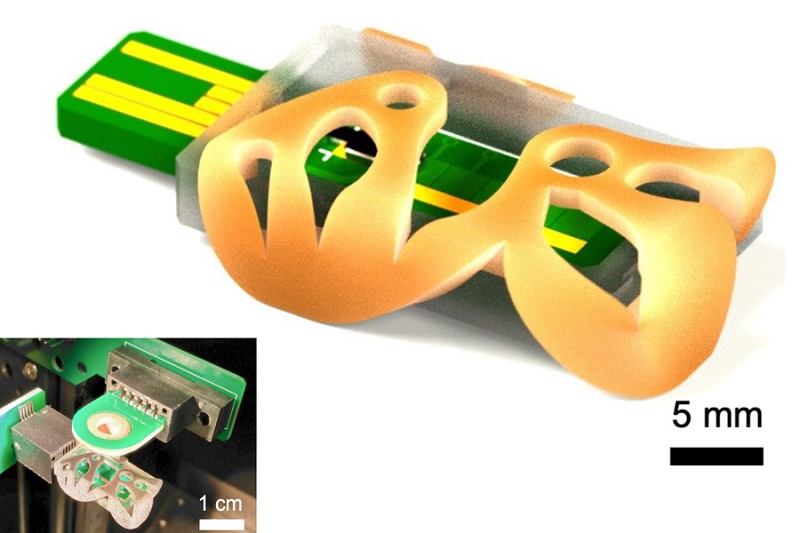
3D Printed Point-Of-Care Mass Spectrometer Outperforms State-Of-The-Art Models
Mass spectrometry is a precise technique for identifying the chemical components of a sample and has significant potential for monitoring chronic illness health states, such as measuring hormone levels... Read more.jpg)
POC Biomedical Test Spins Water Droplet Using Sound Waves for Cancer Detection
Exosomes, tiny cellular bioparticles carrying a specific set of proteins, lipids, and genetic materials, play a crucial role in cell communication and hold promise for non-invasive diagnostics.... Read more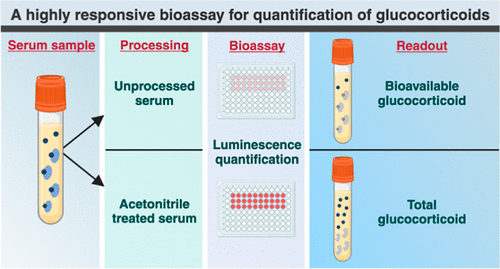
Highly Reliable Cell-Based Assay Enables Accurate Diagnosis of Endocrine Diseases
The conventional methods for measuring free cortisol, the body's stress hormone, from blood or saliva are quite demanding and require sample processing. The most common method, therefore, involves collecting... Read moreHematology
view channel
Next Generation Instrument Screens for Hemoglobin Disorders in Newborns
Hemoglobinopathies, the most widespread inherited conditions globally, affect about 7% of the population as carriers, with 2.7% of newborns being born with these conditions. The spectrum of clinical manifestations... Read more
First 4-in-1 Nucleic Acid Test for Arbovirus Screening to Reduce Risk of Transfusion-Transmitted Infections
Arboviruses represent an emerging global health threat, exacerbated by climate change and increased international travel that is facilitating their spread across new regions. Chikungunya, dengue, West... Read more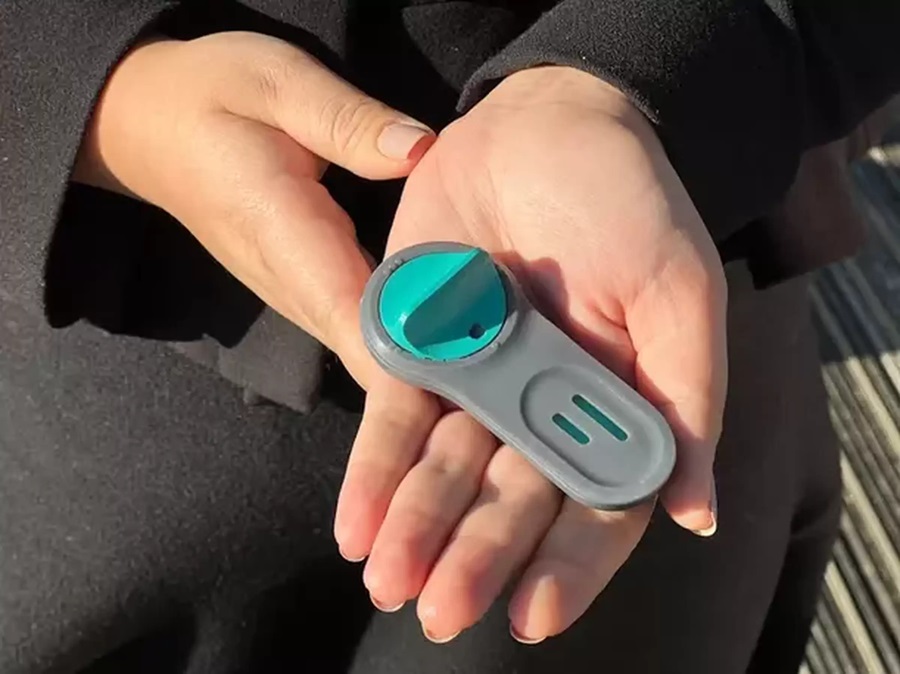
POC Finger-Prick Blood Test Determines Risk of Neutropenic Sepsis in Patients Undergoing Chemotherapy
Neutropenia, a decrease in neutrophils (a type of white blood cell crucial for fighting infections), is a frequent side effect of certain cancer treatments. This condition elevates the risk of infections,... Read more
First Affordable and Rapid Test for Beta Thalassemia Demonstrates 99% Diagnostic Accuracy
Hemoglobin disorders rank as some of the most prevalent monogenic diseases globally. Among various hemoglobin disorders, beta thalassemia, a hereditary blood disorder, affects about 1.5% of the world's... Read moreImmunology
view channel.jpg)
AI Predicts Tumor-Killing Cells with High Accuracy
Cellular immunotherapy involves extracting immune cells from a patient's tumor, potentially enhancing their cancer-fighting capabilities through engineering, and then expanding and reintroducing them into the body.... Read more
Diagnostic Blood Test for Cellular Rejection after Organ Transplant Could Replace Surgical Biopsies
Transplanted organs constantly face the risk of being rejected by the recipient's immune system which differentiates self from non-self using T cells and B cells. T cells are commonly associated with acute... Read more
AI Tool Precisely Matches Cancer Drugs to Patients Using Information from Each Tumor Cell
Current strategies for matching cancer patients with specific treatments often depend on bulk sequencing of tumor DNA and RNA, which provides an average profile from all cells within a tumor sample.... Read more
Genetic Testing Combined With Personalized Drug Screening On Tumor Samples to Revolutionize Cancer Treatment
Cancer treatment typically adheres to a standard of care—established, statistically validated regimens that are effective for the majority of patients. However, the disease’s inherent variability means... Read moreMicrobiology
view channel
Integrated Solution Ushers New Era of Automated Tuberculosis Testing
Tuberculosis (TB) is responsible for 1.3 million deaths every year, positioning it as one of the top killers globally due to a single infectious agent. In 2022, around 10.6 million people were diagnosed... Read more
Automated Sepsis Test System Enables Rapid Diagnosis for Patients with Severe Bloodstream Infections
Sepsis affects up to 50 million people globally each year, with bacteraemia, formerly known as blood poisoning, being a major cause. In the United States alone, approximately two million individuals are... Read moreEnhanced Rapid Syndromic Molecular Diagnostic Solution Detects Broad Range of Infectious Diseases
GenMark Diagnostics (Carlsbad, CA, USA), a member of the Roche Group (Basel, Switzerland), has rebranded its ePlex® system as the cobas eplex system. This rebranding under the globally renowned cobas name... Read more
Clinical Decision Support Software a Game-Changer in Antimicrobial Resistance Battle
Antimicrobial resistance (AMR) is a serious global public health concern that claims millions of lives every year. It primarily results from the inappropriate and excessive use of antibiotics, which reduces... Read morePathology
view channel
New AI Tool Classifies Brain Tumors More Quickly and Accurately
Precision in diagnosing and categorizing tumors is essential for delivering effective treatment to patients. Currently, the gold standard for identifying various types of brain tumors involves DNA methylation-based... Read more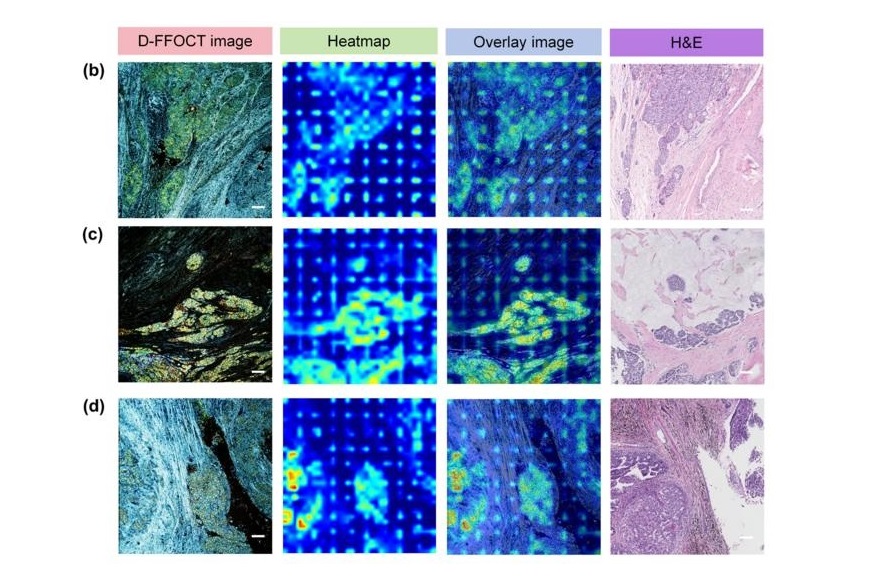
AI Integrated With Optical Imaging Technology Enables Rapid Intraoperative Diagnosis
Rapid and accurate intraoperative diagnosis is essential for tumor surgery as it guides surgical decisions with precision. Traditional intraoperative assessments, such as frozen sections based on H&E... Read more
HPV Self-Collection Solution Improves Access to Cervical Cancer Testing
Annually, over 604,000 women across the world are diagnosed with cervical cancer, and about 342,000 die from this disease, which is preventable and primarily caused by the Human Papillomavirus (HPV).... Read moreHyperspectral Dark-Field Microscopy Enables Rapid and Accurate Identification of Cancerous Tissues
Breast cancer remains a major cause of cancer-related mortality among women. Breast-conserving surgery (BCS), also known as lumpectomy, is the removal of the cancerous lump and a small margin of surrounding tissue.... Read moreTechnology
view channel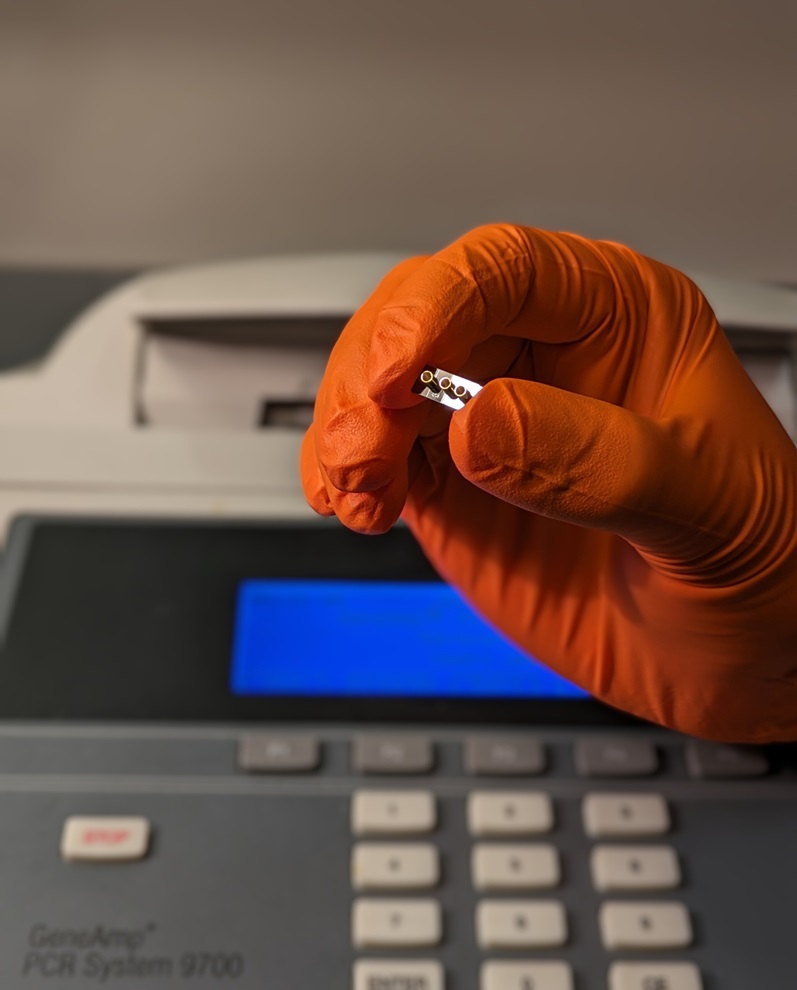
New Diagnostic System Achieves PCR Testing Accuracy
While PCR tests are the gold standard of accuracy for virology testing, they come with limitations such as complexity, the need for skilled lab operators, and longer result times. They also require complex... Read more
DNA Biosensor Enables Early Diagnosis of Cervical Cancer
Molybdenum disulfide (MoS2), recognized for its potential to form two-dimensional nanosheets like graphene, is a material that's increasingly catching the eye of the scientific community.... Read more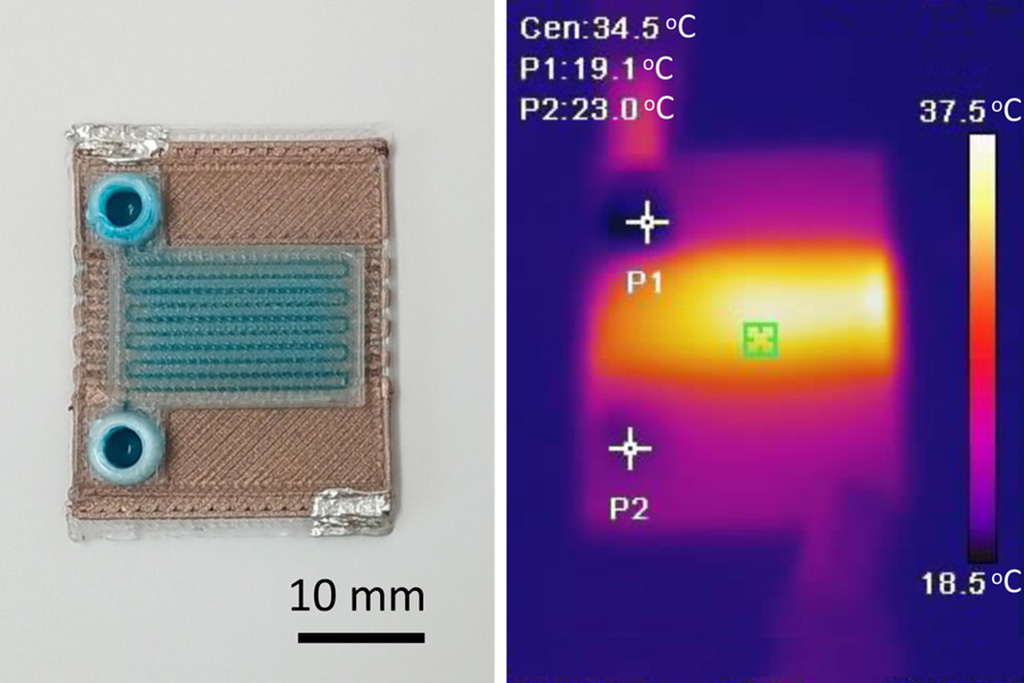
Self-Heating Microfluidic Devices Can Detect Diseases in Tiny Blood or Fluid Samples
Microfluidics, which are miniature devices that control the flow of liquids and facilitate chemical reactions, play a key role in disease detection from small samples of blood or other fluids.... Read more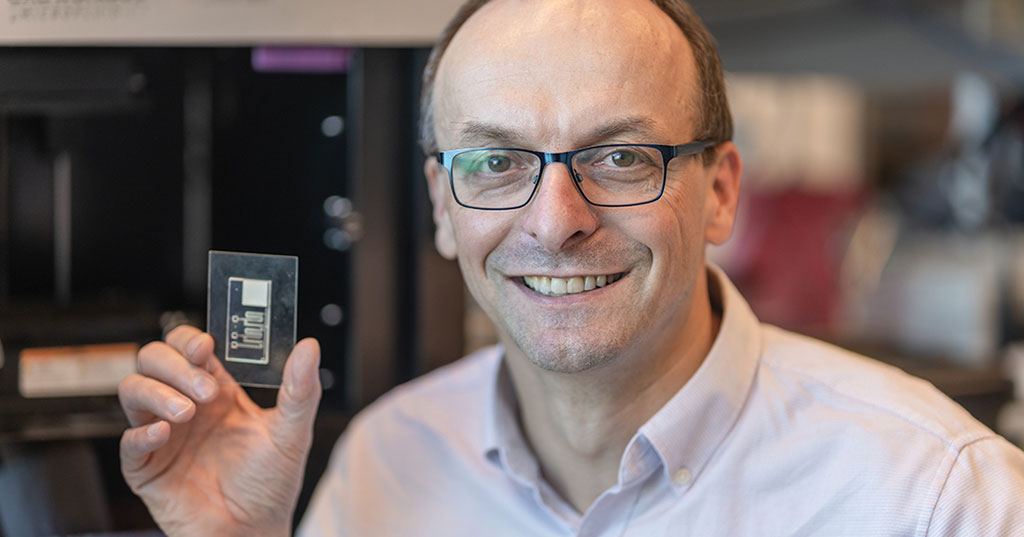
Breakthrough in Diagnostic Technology Could Make On-The-Spot Testing Widely Accessible
Home testing gained significant importance during the COVID-19 pandemic, yet the availability of rapid tests is limited, and most of them can only drive one liquid across the strip, leading to continued... Read moreIndustry
view channel
Danaher and Johns Hopkins University Collaborate to Improve Neurological Diagnosis
Unlike severe traumatic brain injury (TBI), mild TBI often does not show clear correlations with abnormalities detected through head computed tomography (CT) scans. Consequently, there is a pressing need... Read more
Beckman Coulter and MeMed Expand Host Immune Response Diagnostics Partnership
Beckman Coulter Diagnostics (Brea, CA, USA) and MeMed BV (Haifa, Israel) have expanded their host immune response diagnostics partnership. Beckman Coulter is now an authorized distributor of the MeMed... Read more_1.jpg)